Phylogenetics: Searching Lab

|
EEB 5349: Phylogenetics |
| This lab explores different search strategies under the parsimony criterion |
Contents
- 1 Getting started
- 2 Download a copy of the data file
- 3 Create a command file
- 4 Delete all taxa except the first five
- 5 Exhaustive parsimony search
- 6 Determine NNI rearrangements
- 7 Perform an NNI search
- 8 Find the most parsimonious tree for all 35 taxa
- 9 Conduct a second search with SPR swapping
- 10 Now conduct a third search with TBR swapping
- 11 Compare NJ with a comparable heuristic search
- 12 Using SplitsTree
Getting started
Log into your account on the cluster (ssh username@bbcsrv3.biotech.uconn.edu). Type the following:
qlogin
This asks the scheduler to find a node (computer) in the cluster that is currently not busy. It should transfer your session from the head node to a different node. The reason we are using qlogin today is that some of the analyses we are going to run take more than a second or two to run. If all of you ran these jobs on the head node, all of us would notice a significant slowdown in response time (as would anyone else trying to use the cluster).
Download a copy of the data file
If you do not have the angio35.nex file, you can recreate it as follows:
curl http://hydrodictyon.eeb.uconn.edu/people/plewis/courses/phylogenetics/data/angio35.nex > angio35.nex
The word curl stands for "Copy URL" and the program curl fetches the contents of the specified web site (which in this case is not really a web site, it is just a data file). Without the trailing "> angio35.nex", you would see the contents of the angio35.nex file simply spill across your screen. That ">" symbol followed by a file name causes the output of curl to be captured and redirected to a file named angio35.nex.
Create a command file
Now start the nano editor (do not specify a file name because you will be creating a new file) and enter the following text, saving the file as run.nex:
#nexus begin paup; log file=output.txt start replace; execute angio35.nex; end;
There are at least a couple of advantages to creating little NEXUS files like run.nex. For now, the only advantage is that executing run.nex automatically starts a log file so that you will have a record of what you did. Later, when you get in the habit of putting commands in paup blocks, you will appreciate the fact that your data are separated from the commands that initiate analyses. I have many times opened a data file, forgetting about the embedded paup block that starts a long search and immediately overwrites my previous output files!
Note that because we used the replace keyword in the log command, the file output.txt will be overwritten without warning if it exists. Yes, this is called living dangerously but saves some frustration if a run must be restarted.
Delete all taxa except the first five
Using this command
delete 6-.
will cause PAUP* to ignore all taxa except Ephedrasinica, Gnetum_gnemJS, WelwitschiaJS, Ginkgo_biloba, and Pinus_ellCH.
Exhaustive parsimony search
Use the alltrees command to conduct an exhaustive search under the parsimony criterion (parsimony is the default optimality criterion).
You should add the delete and alltrees file to the paup block in your run.nex file, which should now look like this:
#nexus begin paup; log file=output.txt start replace; execute angio35.nex; delete 6-.; alltrees; end;
Start paup, specifying run.nex as the file to execute:
paup run.nex
This analysis should go fast because you now have only 5 taxa.
Determine NNI rearrangements
Because we performed an exhaustive enumeration, we now know which tree is the globally most parsimonious tree. We are thus guaranteed to never find a better tree were we to start an heuristic search with this tree. Let's do an experiment: perform an NNI heuristic search, starting with the best tree, and have PAUP* save all the trees it encounters in this search. In the end, PAUP* will have 5 trees in memory: the starting tree and the 4 trees corresponding to all possible NNI rearrangements of that starting tree.
Before you start the NNI search, use the showtree command to show you the tree obtained from the exhaustive enumeration. Draw this tree on a piece of paper and then draw the 4 possible NNI rearrangements (refer to the description of NNI in my lecture slides if you've forgotten).
Perform an NNI search
Add an hsearch and describe command to your run.nex file, which should afterwards look like this:
#nexus begin paup; log file=output.txt start replace; execute angio35.nex; delete 6-.; alltrees; hsearch start=1 swap=nni nbest=15; describe all; end;
The hsearch command is broken down as follows:
- start=1 starts the search from the tree currently in memory (i.e., the best tree resulting from your exhaustive search using the parsimony criterion)
- swap=nni causes the Nearest-Neighbor Interchange (NNI) method to be used for branch swapping
- nbest=15 saves the 15 best trees found during the search. Thus, were PAUP to examine every possible tree, we would end up saving all of them in memory. The reason this command is needed is that PAUP ordinarily does not save trees that are worse than the best one it has seen thus far. Here, we are interested in seeing the trees that are examined during the course of the search, even if they are not as good as the starting tree.
The describe all command plots the 5 trees currently in memory. The reason we are using the describe command rather than the showtrees command is because we want PAUP to show us the numbers it has assigned to the internal nodes, something that showtrees doesn't do.
Find the most parsimonious tree for all 35 taxa
Modify your run.nex file to conduct a heuristic search on all 35 taxa having the following characteristics:
- The starting trees are each generated by the stepwise addition method, using random addition of sequences (you will employ the addseq and start keywords for this)
- Swap using NNI branch swapping (you will employ the swap keyword for this)
- Reset the nbest option to all because we want to be saving just the best trees, not suboptimal trees (yes, this option is a little confusing).
- Set the random number seed to 5555 using the rseed option (this determines the sequence of pseudorandom numbers used for the random additions; ordinarily you would not need to set the random number seed, but we will do this here to ensure that we all get the same results)
- Do 500 replicate searches; each replicate represents an independent search starting from a different random-addition tree (you will use the nreps keyword for this).
Use the following command to get PAUP to list the options for hsearch:
hsearch ?
Remember you can comment out portions of your Nexus file if you don't want to lose them: e.g.,
#nexus begin paup; log file=output.txt start replace; execute angio35.nex; [delete 6-.;] [alltrees;] hsearch [put your new options here]; [describe all;] end;
Conduct a second search with SPR swapping
Construct another heuristic search using SPR branch swapping. Be sure to reset the random number seed to 5555.
Now conduct a third search with TBR swapping
Wondering about this warning? Multiple hits on islands of unsaved trees may in fact represent different islands When PAUP encounters a new island, it will find all trees composing that particular island in the process of branch swapping. If, in a new search, it encounters any trees already stored in memory, it knows that it has hit an island that it found previously. Note that it would be pointless to continue on this tack, because it will only find all the trees on that island again. For trees retained in memory, PAUP can keep track of which island they belong to (remember that it is possible for trees with the same parsimony score to be in different tree islands!). But for trees that are not retained in memory, PAUP only knows that it has encountered an island of trees having score X; it has no way of finding out how many islands are actually represented amongst the trees having score X.
Compare NJ with a comparable heuristic search
In this exercise, you will conduct a Neighbor-joining (NJ) analysis using HKY distances and compare this with an heuristic search using the minimum evolution criterion. The goal of this section is to demonstrate that it is possible for heuristic searching to find a better tree than NJ, even using the same optimality criterion.
Please quit PAUP* (if it is still running) and start it again. The purpose of restarting is to return all settings to their default values.
Create a new file using nano containing the following lines. Note that we are again using the angio35.nex file:
#nexus begin paup; execute angio35.nex; dset distance=hky objective=me; nj; dscores 1; [!*** NJ score above ***] hsearch start=1; dscores 1; [!*** Heuristic search score above ***] end;
(The text surrounded by square brackets is a comment, and the initial exclamation point ! tells PAUP that you would like this comment to appear in the output.) Run this file in paup by typing the following at the linux prompt:
paup <filename>
(Of course, replace <filename> with the actual name of the file you just created.) Normally, you would not run paup on the cluster directly like this, but this run finishes almost instantly and thus no one will mind. I will show you in a later lab how you should set up more extensive analyses that take a long time to finish.
Once you have figured out what is going on (ask us for help if you are stumped), fix your paup block and re-execute the file. You may need to get PAUP to help you with the criterion setting; type the following to get PAUP to spit out the current settings, then look for criterion near the top of the list:
set ?
In your reanalysis, you should find that the heuristic search starting with the NJ tree found a better tree using the same optimality criterion (minimum evolution) being used by NJ. Neighbor-joining is very popular, but you should be wary of NJ trees involving large numbers of taxa. This analysis involved 35 taxa; for problem of this size or larger, it is almost certain that NJ will not find the best tree.
Using SplitsTree
We will now abandon PAUP* and explore a program called SplitsTree. You will use SplitsTree on your own computer, not the cluster. Go to the SplitsTree web site and download and install a version appropriate for your computer.
Start SplitsTree by double-clicking its icon. After it loads, use Open... from the File menu to navigate to the Examples folder and open bees.nex. Note the webbiness, which indicates that some characters prefer one tree, and other characters prefer a different tree. You can make the tree bigger using your scroll wheel (or the buttons at the top of the main window), and if you select a node, the program allows you to swivel the edges of the graph.
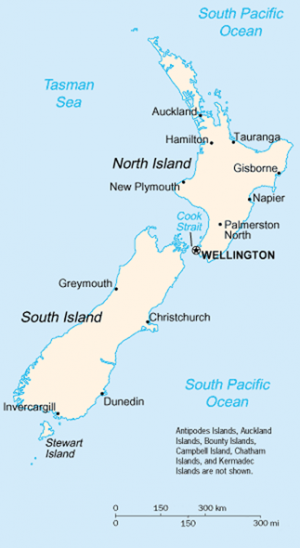
Most phylogeny programs will not make it clear when a tree is not the best way to represent the data. SplitsTree excels at this. Open the example file colors.nex. The distance matrix provided in this case comprises pairwise distances between different colors, which you would not expect to be especially treelike.The example file south.nex contains the pairwise distances between cities on the south island of New Zealand (plus Wellington, which is the southernmost city on the north island). A quick look at the map of New Zealand on the right will convince you that the "tree" shown by SplitsTree is actually a fairly good map of the south island of New Zealand!
SplitsTree makes use of the NEXUS format, so you can create a distance matrix in PAUP* and then read it in to SplitsTree. SplitsTree uses several types of blocks that PAUP* does not recognize, however, including Splits, st_graph and st_assumptions. Some of these blocks are added to your data file when you execute it, which is why when you try to close a window SplitsTree asks you whether you want to save the modified data file.
Return to the bees.nex example. To show the split weights on the tree, choose Select Edges from the Edit menu, then choose Format Nodes and Edges... from the View menu. In the dialog box that appears, check the Weights checkbox beside Edge labels.
The SplitsTree program allows you to save the tree in various graphics file formats, including jpg, eps, svg, gif or png. Use the Export Image... menu command from the File menu to create a file of one of these types.