Phylogenetics: Compositional Heterogeneity Lab
From EEBedia
Revision as of 19:44, 30 March 2014 by Paul Lewis (Talk | contribs) (→Simulated data and the true tree)

|
EEB 5349: Phylogenetics |
| The goal of this lab is to introduce you to the influence of compositional heterogeneity on phylogeny. Compositional heterogeneity means that the equilibrium nucleotide frequencies (or amino acid frequencies for protein data) change across the tree, something that is not accounted for by the standard nucleotide and amino acid models, which assume stationarity (transition probabilities do not change across the tree and one set of equilibrium frequencies applies to every point along any edge of the tree). Non-stationarity can lead to compositional attraction artifacts in which tips with similar nucleotide composition group together even though they may be completely unrelated. |
Contents
Under Constructions (should be finished later today, March 30, 2014)
Simulated data and the true tree
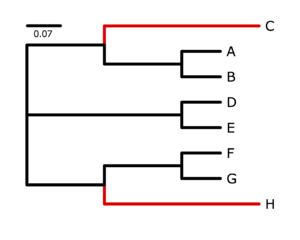
The tree on the right was used to simulate data using p4. Peter Foster's program p4 specializes in simulating and analyzing data in which nucleotide composition varies across the tree. Here, we will use data simulated using p4, but will perform analyses using a different program, nhPhyloBayes, written by Samuel Blanquart.Obtaining nhPhyloBayes
nhPhyloBayes is available as a tar archive,
Ordinary models yield the compositional attraction tree
Non-homogeneous models yield the true tree
- Question answer