Difference between revisions of "Phylogenetics: Compositional Heterogeneity Lab"
Paul Lewis (Talk | contribs) (→Ordinary models yield the compositional attraction tree) |
Paul Lewis (Talk | contribs) (→Ordinary models yield the compositional attraction tree) |
||
| Line 63: | Line 63: | ||
* ''In what file will the globally optimum tree be saved?'' {{title|gtrig-alltrees.tre|answer}} | * ''In what file will the globally optimum tree be saved?'' {{title|gtrig-alltrees.tre|answer}} | ||
</div> | </div> | ||
| + | Now run the <tt>paup.nex</tt> file in PAUP*. You can start it like this using qsub: | ||
| + | qsub paup paup.nex | ||
== Non-homogeneous models yield the true tree == | == Non-homogeneous models yield the true tree == | ||
Revision as of 22:56, 30 March 2014

|
EEB 5349: Phylogenetics |
| The goal of this lab is to introduce you to the influence of compositional heterogeneity on phylogeny. |
Compositional heterogeneity means that the equilibrium nucleotide frequencies (or amino acid frequencies for protein data) change across the tree, something that is not accounted for by the standard nucleotide and amino acid models, which assume stationarity (transition probabilities do not change across the tree and one set of equilibrium frequencies applies to every point along any edge of the tree). Non-stationarity can lead to compositional attraction artifacts in which tips with similar nucleotide composition group together even though they may be completely unrelated
Contents
Under Construction (should be finished later today, March 30, 2014)
Simulated data and the true tree
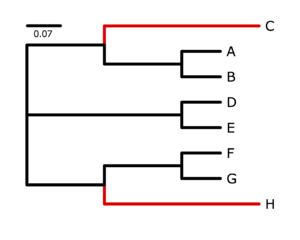
The program p4, written by Peter Foster, specializes in simulating and analyzing data in which nucleotide composition varies across the tree. I used p4 to simulate data on the tree show on the right. The black-colored lineages were characterized by an AT-biased nucleotide composition very different from the red-colored lineages, which were strongly GC-biased. My goal was to generate a data set that would be very susceptible to nucleotide compositional attraction under ordinary substitution models, and in that I succeeded (as you will see). Taxa C and H share many similarities due to the large number of G and C bases they have independently acquired from their AT-rich ancestors, and it will be very tempting for an ordinary nucleotide model such as GTR to place C and H together.First, log in to the cluster and use qlogin to acquire a free slot. Then download the 500-site simulated data set to a directory named nhlab in your home directory on the cluster:
mkdir nhlab cd nhlab curl http://hydrodictyon.eeb.uconn.edu/people/plewis/courses/phylogenetics/labs/sim500.nex > sim500.nex
At the end of this lab, I will show you how I simulated this data set in p4, but I don't want you to take time doing that now - the analysis will take quite a bit of lab time so I want you to get started on that right away.
Running nhPhyloBayes
You will perform analyses using the program nhPhyloBayes, written by Samuel Blanquart. nhPhyloBayes is available as a tar archive from nh_PhyloBayes_0.2.3.tar, but I have already downloaded, compiled, and installed nhPhyloBayes for you on the cluster, so you can start your run right away. The following command assumes you are currently inside ~/nhlab, where ~ is a symbol that means your home directory. Note that we are not using qsub to start this run; I'll explain everything in a minute, but for now just (very carefully) type in the following line (or, better yet, copy and paste it) and then press the enter key:
/common/nh/nhpb -d sim500.nex -f sim500 -m bp -x ~/nhlab -y /common/nh/ -Q
Here is a complete explanation of everything you've accomplished on that one line:
- /common/nh/nhpb starts the nhPhyloBayes program. It is located in the /common/nh directory, which is probably not in the path that the system searches to find programs, so you must be explicit and provide the full path
- -d sim500.nex specifies the name of the data file, which should be located in the directory you are in when you issue this command
- -f sim500 specifies the prefix that will be used for all output files (the fact that it is the same as the data file name prefix just illustrates my lack of imagination)
- -m bp specifies the model. nhPhyloBayes does not give you a lot of choices with respect to the model, and bp is the simplest nh (non-homogeneous) model offered by the program
- -x ~/nhlab should be the name of the directory you are in when you issue the command (note: no trailing slash)
- -y /common/nh/ should be the name of the directory where the nhPhyloBayes program is located (note: this time you do add a trailing slash)
- -Q tells nhPhyloBayes to start your program using qsub. So, even though it looked like we avoided using qsub, in reality we used qsub to start the analysis
You should be able to check whether the program is running using the qstat command. If the program did not start, let one of us know and we'll help you get it going. Once you get the program going, we will let it run for about an hour before checking the results.
Ordinary models yield the compositional attraction tree
Before seeing how nhPhyloBayes does with this example, I want you to convince yourself that ordinary models are severely mislead by the strong convergence in nucleotide composition exhibited by these data. Create a nexus file named paup.nex inside your ~/nhlab directory with the following contents:
#nexus begin paup; log file=pauplog.txt start replace; set crit=like; exe sim500.nex; lset nst=6 rmatrix=estimate basefreq=estimate rates=gamma shape=estimate pinvar=estimate; nj; lscores 1; lset rmatrix=previous basefreq=previous shape=previous pinvar=previous; alltrees; savetrees file=gtrig-alltrees.tre brlens; log stop; end;
Answer these questions about this file before you try to run it in PAUP*:
- What will be the name of the file in which PAUP's output will be stored? answer
- What is the optimality criterion that will be used for the search? answer
- Why is a neighbor joining tree generated? answer
- What model is used? answer
- What is the purpose of the second lset command? answer
- What type of search will be conducted? answer
- How many tree topologies will PAUP examine? answer
- In what file will the globally optimum tree be saved? answer
Now run the paup.nex file in PAUP*. You can start it like this using qsub:
qsub paup paup.nex
Non-homogeneous models yield the true tree
- Question answer