Difference between revisions of "Phylogenetics: BayesTraits Lab"
Paul Lewis (Talk | contribs) |
Paul Lewis (Talk | contribs) (→Estimating ancestral states) |
||
| (93 intermediate revisions by 3 users not shown) | |||
| Line 4: | Line 4: | ||
|<span style="font-size: x-large">[http://hydrodictyon.eeb.uconn.edu/eebedia/index.php/Phylogenetics:_Syllabus EEB 349: Phylogenetics]</span> | |<span style="font-size: x-large">[http://hydrodictyon.eeb.uconn.edu/eebedia/index.php/Phylogenetics:_Syllabus EEB 349: Phylogenetics]</span> | ||
|- | |- | ||
| − | |In this lab you will learn how to use the program BayesTraits, written by Andrew Meade and Mark Pagel. BayesTraits can perform several analyses related to evaluating evolutionary correlation in discrete morphological traits | + | |In this lab you will learn how to use the program BayesTraits, written by Andrew Meade and Mark Pagel. BayesTraits can perform several analyses related to evaluating evolutionary correlation and ancestral state reconstruction in discrete morphological traits. |
|} | |} | ||
| − | + | == Download BayesTraits == | |
| − | + | Login to Xanadu and request a machine as usual: | |
| − | = | + | srun --pty -p mcbstudent --qos=mcbstudent bash |
| − | Download BayesTraits from [http://www.evolution. | + | Download BayesTraits from [http://www.evolution.rdg.ac.uk/BayesTraitsV3.0.2/BayesTraitsV3.0.2.html Mark Pagel's web site] using curl. You can get the tar archive linked to the web site as "BayesTraits V3.0.2 - Linux 64" onto Xanadu however you like, but I think the easiest way is to just use curl: |
| − | === Download the tree and data files | + | curl -O http://www.evolution.rdg.ac.uk/BayesTraitsV3.0.2/Files/BayesTraitsV3.0.2-Linux.tar.gz |
| + | |||
| + | Now unpack the gzipped "tape archive" as follows: | ||
| + | |||
| + | tar zxvf BayesTraitsV3.0.2-Linux.tar.gz | ||
| + | |||
| + | This will create a directory named BayesTraitsV3.0.2-Linux. The BayesTraitsV3.0.2-Linux folder contains the program itself along with several tree and data files (e.g. <tt>Primates.txt</tt> and <tt>Primates.trees</tt>). I will hereafter refer to the folder containing these files as simply the '''BayesTraits folder'''. Go back to Mark Pagel's web site and '''download the manual''' for BayesTraits. This is a PDF file and should open in your browser window. | ||
| + | |||
| + | === A little aside on tar files === | ||
| + | Data used to be stored on magnetic tape, not hard drives, and the tar (tape archive) program is what was used to move files to and from the tape. This tells you something about how old the tar format is because perhaps none of you have even seen a magnetic tape used for data storage! The tar command takes all the files in a directory and simply concatenates them into one gigantic file. It also preserves file permissions and the directory structure. The four letters after the command name tar are zxvf. These stand for the following: | ||
| + | * z = uncompress (the gz at the end of the file tells you it is a compressed archive, so the z tells tar to uncompress it before unpacking it) | ||
| + | * x = extract (unpack the archive into individual files. You would use c here if you were creating a tar file) | ||
| + | * v = verbose (tell us what's going on as you unpack) | ||
| + | * f = file (this tells tar that the file name is coming next, so don't put f earlier in the list) | ||
| + | |||
| + | This tar file has been compressed using the program gzip, which adds the gz ending to the file name. Most tar files are compresses with gzip or some similar algorithm so that the file requires less time to move across the internet. | ||
| + | |||
| + | == Download the tree and data files == | ||
For this exercise, you will use data and trees used in the SIMMAP analyses presented in this paper (you should recognize the names of at least two of the authors of this paper): | For this exercise, you will use data and trees used in the SIMMAP analyses presented in this paper (you should recognize the names of at least two of the authors of this paper): | ||
| − | Jones C.S., Bakker F.T., Schlichting C.D., Nicotra A.B. 2009. Leaf shape evolution in the South African genus Pelargonium L'Her. (Geraniaceae). Evolution. 63:479–497. | + | <blockquote>Jones C.S., Bakker F.T., Schlichting C.D., Nicotra A.B. 2009. Leaf shape evolution in the South African genus ''Pelargonium'' L'Her. (Geraniaceae). Evolution. 63:479–497.</blockquote> |
| − | The data and trees were not made available in the online supplementary materials for this paper, but I have obtained permission to use them for this laboratory exercise. The links below are password-protected, so ask us for the username and password before clicking on the links: | + | The data and trees were not made available in the online supplementary materials for this paper, but I have obtained permission to use them for this laboratory exercise. |
| + | <!-- The links below are password-protected, so ask us for the username and password before clicking on the links: --> | ||
| − | :[http://hydrodictyon.eeb.uconn.edu/people/plewis/courses/phylogenetics/ | + | :[http://hydrodictyon.eeb.uconn.edu/people/plewis/courses/phylogenetics/data/pelly.txt pelly.txt] This is the data file. It contains data for two traits (see below) for 154 taxa in the plant genus ''Pelargonium''. |
| − | :[http://hydrodictyon.eeb.uconn.edu/people/plewis/courses/phylogenetics/ | + | :[http://hydrodictyon.eeb.uconn.edu/people/plewis/courses/phylogenetics/data/pelly.tre pelly.tre] This is the tree file. It contains 99 trees sampled from an MCMC analysis of DNA sequences. |
| − | === | + | <!-- |
| − | + | If you are using version 1 of BayesTraits, it will complain about basal polytomies in the trees. The following versions of the files provide a workaround (I deleted one taxon from both the data file and the tree file to eliminate the basal polytomy): | |
| + | :[http://hydrodictyon.eeb.uconn.edu/people/plewis/courses/phylogenetics/restricted/pellyrooted.txt pellyrooted.txt] This is the data file. It contains data for two traits (see below) for 154 taxa in the plant genus ''Pelargonium''. | ||
| + | :[http://hydrodictyon.eeb.uconn.edu/people/plewis/courses/phylogenetics/restricted/pellyrooted.tre pellyrooted.tre] This is the tree file. It contains 99 trees sampled from an MCMC analysis of DNA sequences. | ||
| + | --> | ||
| + | You should move these files to a new folder that you create for this lab. For example | ||
| + | cd # cd alone returns you to your home directory | ||
| + | mkdir pelly | ||
| + | cd pelly | ||
| + | curl -O http://hydrodictyon.eeb.uconn.edu/people/plewis/courses/phylogenetics/data/pelly.txt | ||
| + | curl -O http://hydrodictyon.eeb.uconn.edu/people/plewis/courses/phylogenetics/data/pelly.tre | ||
| + | |||
| + | == Assessing the strength of association between two binary characters == | ||
| + | |||
| + | The first thing we will do is see if the two characters (leaf dissection and leaf venation) in <tt>pelly.txt</tt> are evolutionarily correlated. | ||
| + | |||
| + | === Trait 1: Leaf dissection === | ||
| + | The '''leaf dissection''' trait comprises two states (I've merged some states in the original data matrix to produce just 2 states): | ||
| + | * 0 means leaves are ''entire'' (''unlobed'' or ''shallowly lobed'' in the original study), and | ||
| + | * 1 means leaves are ''dissected'' (''lobed'', ''deeply lobed'', or ''dissected'' in the original study). | ||
| + | |||
| + | === Trait 2: Leaf venation === | ||
| + | The '''leaf venation''' trait comprises two states: | ||
| + | * 0 means leaves are ''pinnately veined'' (one main vein runs down the long axis of the leaf blade), and | ||
| + | * 1 means leaves are ''palmately veined'' (several major veins meet at the base of the leaf). | ||
| + | |||
| + | To test whether these two traits are correlated, we will estimate the '''marginal likelihood''' under two models. The independence model assumes that the two traits are uncorrelated. The dependence model allows the two traits to be correlated in their evolution. The model with the higher marginal likelihood will be the preferred model. You will recall that we discussed both of these models in lecture, and also discussed the '''stepping-stone method''' that BayesTraits uses to evaluate models. You may wish to pull up those lectures to help answer the questions that you will encounter momentarily, as well as the BayesTraits manual. | ||
| + | |||
| + | === Maximum Likelihood: Independence model === | ||
| + | |||
| + | Type the following to start the BayesTraits program (assuming you are in the pelly folder and that the BayesTraitsV3.0.2-Linux is a "sister" folder: | ||
| + | ../BayesTraitsV3.0.2-Linux/BayesTraitsV3 pelly.tre pelly.txt | ||
| + | |||
| + | You should see this selection appear: | ||
| + | Please select the model of evolution to use. | ||
| + | 1) MultiState | ||
| + | 2) Discrete: Independent | ||
| + | 3) Discrete: Dependant | ||
| + | 4) Continuous: Random Walk (Model A) | ||
| + | 5) Continuous: Directional (Model B) | ||
| + | 6) Continuous: Regression | ||
| + | 7) Independent Contrast | ||
| + | 8) Independent Contrast: Correlation | ||
| + | 9) Independent Contrast: Regression | ||
| + | 10) Discrete: Covarion | ||
| + | Press the 2 key and hit enter to select the Independent model. Now you should see these choices appear: | ||
| + | Please Select the analysis method to use. | ||
| + | 1) Maximum Likelihood. | ||
| + | 2) MCMC | ||
| + | Press the 1 key and hit enter to select maximum likelihood. Now you should see some output showing the choices you explicitly (or implicitly) made: | ||
| + | Options: | ||
| + | Model: Discete Independant | ||
| + | Tree File Name: pelly.tre | ||
| + | Data File Name: pelly.txt | ||
| + | Log File Name: pelly.txt.log.txt | ||
| + | Save Initial Trees: False | ||
| + | Save Trees: False | ||
| + | Summary: False | ||
| + | Seed 3162959925 | ||
| + | Analsis Type: Maximum Likelihood | ||
| + | ML attempt per tree: 10 | ||
| + | ML Max Evaluations: 20000 | ||
| + | ML Tolerance: 0.000001 | ||
| + | ML Algorithm: BOBYQA | ||
| + | Rate Range: 0.000000 - 100.000000 | ||
| + | Precision: 64 bits | ||
| + | Cores: 1 | ||
| + | No of Rates: 4 | ||
| + | Base frequency (PI's) None | ||
| + | Character Symbols: 00,01,10,11 | ||
| + | Using a covarion model: False | ||
| + | Restrictions: | ||
| + | alpha1 None | ||
| + | beta1 None | ||
| + | alpha2 None | ||
| + | beta2 None | ||
| + | Tree Information | ||
| + | Trees: 99 | ||
| + | Taxa: 154 | ||
| + | Sites: 1 | ||
| + | States: 4 | ||
| + | Now type <tt>run</tt> and hit enter to perform the analysis, which will consist of estimating the parameters of the independent model on each of the 99 trees contained in the pelly.tre file. | ||
| + | Tree No Lh alpha1 beta1 alpha2 beta2 Root - P(0,0) Root - P(0,1) Root - P(1,0) Root - P(1,1) | ||
| + | 1 -157.362972 53.767527 34.523176 35.319157 20.707416 0.249998 0.250002 0.249998 0.250002 | ||
| + | 2 -158.179984 53.313539 34.182683 36.038859 20.997536 0.249999 0.250001 0.249999 0.250001 | ||
| + | . | ||
| + | . | ||
| + | . | ||
| + | 98 -156.647307 52.357626 36.749282 27.270771 13.086248 0.250244 0.249756 0.250244 0.249756 | ||
| + | 99 -156.532925 52.321467 36.641688 27.402067 13.200124 0.250234 0.249767 0.250233 0.249766 | ||
| + | You will notice that BayesTraits created a new file: <tt>pelly.txt.Log.txt</tt>. '''Rename this file''' <tt>ml-independent.txt</tt> so that it will not be overwritten the next time you run BayesTraits: | ||
| + | mv pelly.txt.Log.txt ml-independent.txt | ||
| + | |||
| + | Try to answer these questions using the output you have generated (you'll need to consult the BayesTraits manual, but ask us if anything doesn't make sense): | ||
<div style="background-color:#ccccff"> | <div style="background-color:#ccccff"> | ||
| − | * | + | * ''Which occurs at a faster rate: pinnate to palmate, or palmate to pinnate?'' {{title|the 0 (pinnate) to 1 (palmate) transition occurs at a faster rate|answer}} |
| − | * | + | * ''Which occurs at a faster rate: entire to dissected, or dissected to entire?'' {{title|the 0 (entire) to 1 (dissected) transition occurs at a faster rate|answer}} |
| − | + | * ''What do you think Root - P(1,1) means (i.e. the last column of numbers)?'' {{title|this is the probability that leaves were both dissected and palmately veined at the root of the tree|answer}} | |
| − | * | + | |
</div> | </div> | ||
| − | === | + | === Maximum Likelihood: Dependence model === |
| − | '' | + | Run BayesTraits again, this time typing 3 on the first screen to choose the dependence model and again typing 1 on the second screen to select maximum likelihood. You should see this output showing the options selected: |
| + | Options: | ||
| + | Model: Discete Dependent | ||
| + | Tree File Name: pelly.tre | ||
| + | Data File Name: pelly.txt | ||
| + | Log File Name: pelly.txt.log.txt | ||
| + | Summary: False | ||
| + | Seed 3601265953 | ||
| + | Analsis Type: Maximum Likelihood | ||
| + | ML attempt per tree: 10 | ||
| + | Precision: 64 bits | ||
| + | Cores: 1 | ||
| + | No of Rates: 8 | ||
| + | Base frequency (PI's) None | ||
| + | Character Symbols: 00,01,10,11 | ||
| + | Using a covarion model: False | ||
| + | Restrictions: | ||
| + | q12 None | ||
| + | q13 None | ||
| + | q21 None | ||
| + | q24 None | ||
| + | q31 None | ||
| + | q34 None | ||
| + | q42 None | ||
| + | q43 None | ||
| + | Tree Information | ||
| + | Trees: 99 | ||
| + | Taxa: 154 | ||
| + | Sites: 1 | ||
| + | States: 4 | ||
| + | Run the analysis. Here is an example of the output produced after you type <tt>run</tt> to start the analysis. The column headers don't quite line up with the columns, but you can fix this in a text editor or by copying and pasting the table-like output from the log file into a spreadsheet program: | ||
| + | Tree No Lh q12 q13 q21 q24 q31 q34 q42 q43 Root - P(0,0) Root - P(0,1) Root - P(1,0) Root - P(1,1) | ||
| + | 1 -151.930254 66.451053 37.783888 0.000000 62.220033 23.997490 23.299393 46.110432 36.632979 0.24999 0.249981 0.250026 0.250000 | ||
| + | 2 -152.925691 67.152271 38.611193 0.000000 60.925185 24.514488 23.937433 45.313366 37.199310 0.24999 0.249983 0.250023 0.250001 | ||
| + | . | ||
| + | . | ||
| + | . | ||
| + | 98 -150.816306 36.534843 27.359325 0.000000 66.563262 19.823546 24.944519 63.940577 31.074092 0.250048 0.249750 0.250304 0.249898 | ||
| + | 99 -150.712705 37.316351 27.260833 0.000000 64.364694 20.107653 25.004246 60.945163 31.658536 0.250030 0.249779 0.250272 0.249919 | ||
| + | '''Before doing anything else, rename the file''' <tt>pelly.txt.Log.txt</tt> to <tt>ml-dependent.txt</tt> so that it will not be overwritten the next time you run BayesTraits. | ||
| − | ''' | + | Try to answer these questions using the output you have generated: |
| − | + | <div style="background-color:#ccccff"> | |
| − | + | * ''What type of joint evolutionary transitions seem to often have very low rates (look for an abundance of zeros in a column)?'' {{title|q21, which involves entire leaves changing from palmate to pinnate, and q43, which involves dissected leaves changing from palmate to pinnate|answer}} | |
| + | * ''What type of joint evolutionary transitions seem to often have very high rates (look for columns with rates in the hundreds)?'' {{title|q12, which involves entire leaves changing from pinnate to palmate, and q13, which involves pinnate leaves changing from entire to dissected|answer}} | ||
| + | </div> | ||
| − | + | === Bayesian MCMC: Dependence model === | |
| − | + | Run BayesTraits again, typing 3 on the first screen to choose the dependence model and this time typing 2 on the second screen to select MCMC. You should see this output showing the options selected: | |
| + | Options: | ||
| + | Model: Discete Dependent | ||
| + | Tree File Name: pelly.tre | ||
| + | Data File Name: pelly.txt | ||
| + | Log File Name: pelly.txt.log.txt | ||
| + | Summary: False | ||
| + | Seed 3792635164 | ||
| + | Precision: 64 bits | ||
| + | Cores: 1 | ||
| + | Analysis Type: MCMC | ||
| + | Sample Period: 1000 | ||
| + | Iterations: 1010000 | ||
| + | Burn in: 10000 | ||
| + | MCMC ML Start: False | ||
| + | Schedule File: pelly.txt.log.txt.Schedule.txt | ||
| + | Rate Dev: AutoTune | ||
| + | No of Rates: 8 | ||
| + | Base frequency (PI's) None | ||
| + | Character Symbols: 00,01,10,11 | ||
| + | Using a covarion model: False | ||
| + | Restrictions: | ||
| + | q12 None | ||
| + | q13 None | ||
| + | q21 None | ||
| + | q24 None | ||
| + | q31 None | ||
| + | q34 None | ||
| + | q42 None | ||
| + | q43 None | ||
| + | Prior Information: | ||
| + | Prior Categories: 100 | ||
| + | q12 uniform 0.00 100.00 | ||
| + | q13 uniform 0.00 100.00 | ||
| + | q21 uniform 0.00 100.00 | ||
| + | q24 uniform 0.00 100.00 | ||
| + | q31 uniform 0.00 100.00 | ||
| + | q34 uniform 0.00 100.00 | ||
| + | q42 uniform 0.00 100.00 | ||
| + | q43 uniform 0.00 100.00 | ||
| + | Tree Information | ||
| + | Trees: 99 | ||
| + | Taxa: 154 | ||
| + | Sites: 1 | ||
| + | States: 4 | ||
| + | '''Before typing run''' type the following command, which tells BayesTraits to change all priors from the default Uniform(0,100) to an Exponential distribution with mean 30: | ||
| + | pa exp 30 | ||
| − | ''' | + | <div style="background-color:#ccccff"> |
| − | + | * ''Why am I suggesting this switch?'' {{title|think about the support of a Uniform(0,100) distribution vs. the support of an Exponential(1/30) distribution|answer}} | |
| + | * ''Why 30?'' {{title|calculate the variance of a Uniform(0,100) distribution vs. the variance of an Exponential(1/30) distribution|answer}} | ||
| + | </div> | ||
| − | ''' | + | Also type the following to ask BayesTraits to perform a stepping-stone analysis: |
| + | stones 100 10000 | ||
| + | Now run the analysis. This will estimate 100 ratios to brook the gap between posterior and prior, using a sample size of 10000 for each "stone". | ||
| + | Here is an example of the output produced after you type <tt>run</tt> to start the analysis: | ||
| + | Iteration Lh Tree No q12 q13 q21 q24 q31 q34 q42 q43 Root - P(0,0) Root - P(0,1) Root - P(1,0) Root - P(1,1) | ||
| + | 11000 -155.195365 78 14.423234 34.800270 8.845985 45.927148 12.622435 50.476188 52.844895 32.149168 0.250068 0.249969 0.249994 0.249968 | ||
| + | 12000 -154.161705 82 64.601017 12.382781 9.259134 51.796365 12.002095 23.744903 30.316089 21.865930 0.249936 0.249957 0.250095 0.250012 . | ||
| + | . | ||
| + | . | ||
| + | 1009000 -154.343996 30 33.555198 50.086092 11.294490 38.518607 24.461032 47.295157 43.477964 21.726938 0.250057 0.249939 0.250045 0.249959 | ||
| + | 1010000 -154.195259 87 29.584898 35.410909 2.003582 61.981073 16.976124 14.895266 49.111354 14.419644 0.251115 0.247854 0.252551 0.248480 | ||
| + | '''Before doing anything else, rename the file''' <tt>pelly.txt.Log.txt</tt> to <tt>mcmc-dependent.txt</tt>, and <tt>pelly.txt.log.Stones.txt</tt> to <tt>mcmc-dependent.Stones.txt</tt> so that they will not be overwritten the next time you run BayesTraits. | ||
| − | + | You will notice a column not present in the likelihood analysis named ''Tree No'' that shows which of the 99 trees in the supplied <tt>pelly.tre</tt> treefile was chosen at random to be used for that particular sample point. BayesTraits is sampling trees from the posterior distribution here; it cannot ''actually'' sample trees from the posterior because we have given it only data for two morphological characters, which would not provide nearly enough information to estimate the phylogeny for 154 taxa. It is as if we had given BayesTraits sequence data as well as our 2 morphological characters and it was using only the sequence data to estimate the posterior distribution of trees and edge lengths and only the morphological data to estimate rates for the morphological characters. | |
| − | + | Try to answer these questions using the output you have generated: | |
| + | <div style="background-color:#ccccff"> | ||
| + | * ''What is the log marginal likelihood estimated using the stepping-stone method? This value is listed on the last line of the file <tt>mcmc-dependent.Stones.txt</tt> (your value may differ from mine slightly)'' {{title|I got -160.567444 |answer}} | ||
| + | </div> | ||
| − | + | === Bayesian MCMC: Independence model === | |
| − | + | Run BayesTraits again, this time specifying the Independent model, and again using MCMC, <tt>pa exp 30</tt>, and <tt>stones 100 10000</tt>. Rename the output file from <tt>pelly.txt.log.txt</tt> to <tt>mcmc-independent.txt</tt>. Also rename <tt>pelly.txt.log.Stones.txt</tt> to <tt>mcmc-independent.Stones.txt</tt>. | |
| − | + | <div style="background-color:#ccccff"> | |
| − | + | * ''What is the estimated log marginal likelihood for this analysis using the stepping-stone method?'' {{title|I got -162.693620|answer}} | |
| − | + | * ''Which is the better model (dependent or independent) according to these estimates of marginal likelihood?'' {{title|the dependent model has a slightly higher marginal likelihood and is thus preferred|answer}} | |
| + | </div> | ||
| − | + | === Bayesian Reversible-jump MCMC === | |
| − | + | ||
| − | + | ||
| − | + | Run BayesTraits again, specifying Dependent model, MCMC and, this time, specify the reversible-jump approach using the command | |
| − | + | rj exp 30 | |
| − | + | The previous command also sets the prior. Type <tt>run</tt> to start, then when it finishes rename the output file <tt>rjmcmc-dependent.txt</tt>. | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | The reversible-jump approach carries out an MCMC analysis in which the number of model parameters (the dimension of the model) potentially changes from one iteration to the next. The full model allows each of the 8 rate parameters to be estimated separately, while other models restrict the values of some rate parameters to equal the values of other rate parameters. The output contains a column titled '''Model string''' that summarizes the model in a string of 8 symbols corresponding to the 8 rate parameters q12, q13, q21, q24, q31, q34, q42, and q43. For example, the model string "'1 0 Z 0 1 1 0 2" sets q21 to zero (Z), q13=q24=q42 (parameter group 0), q12=q31=q34 (parameter group 1), and q43 has its own non-zero value distinct from parameter groups 0 and 1. | |
| − | + | ||
| − | + | You could copy the "spreadsheet" part of the output file into Excel and sort by the model string column, but let's instead use Python to summarize the output file. Download (e.g. using curl) the file [http://hydrodictyon.eeb.uconn.edu/people/plewis/courses/phylogenetics/data/btsummary.py btsummary.py] file and run it as follows: | |
| + | python btsummary.py | ||
| + | This should produce counts of model strings. (If it doesn't, check to make sure your output file is named <tt>rjmcmc-dependent.txt</tt> because ''btsummary.py'' tries to open a file by that name.) Answer the following questions using the counts provided by ''btsummary.py''. | ||
| + | <div style="background-color:#ccccff"> | ||
| + | * ''Which model string is most common?'' {{title|I got 0 0 Z 0 0 0 0 0 with count 979|answer}} | ||
| + | * ''What does this model imply?'' {{title|all rates are the same except q21, which is forced to have rate zero. q21 equals 0 implies that entire,palmate leaves never change to entire,pinnate|answer}} | ||
| + | </div> | ||
| − | + | Notice that many (but not all) model strings have Z for q21. One way to estimate the marginal posterior probability of the hypothesis that q21=0 is to sum the counts for all model strings that have Z in that third position corresponding to q21. While it is pretty easy to add these numbers in your head, let's modify ''btsummary.py'' to do this for us (this might come in useful if you ever encounter results that are more complex): open ''btsummary.py'' and locate the line containing the [https://en.wikipedia.org/wiki/Regular_expression regular expression] search that pulls out all the model strings from the BayesTrait output file: | |
| − | # | + | model_list = re.findall("'[Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9]", stuff, re.M | re.S) |
| − | + | The <tt>re.findall</tt> function performs a regular expression search of the text stored in the variable stuff looking for strings that have a series of 8 space-separated characters, each of which is ''either'' the character Z ''or'' a digit between 0 and 9 (inclusive). Copy this line, then comment out one copy by starting the line with the hash (#) character: | |
| − | # | + | #model_list = re.findall("'[Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9]", stuff, re.M | re.S) |
| − | + | model_list = re.findall("'[Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9]", stuff, re.M | re.S) | |
| − | + | Now modify the copy such that it counts only models with Z in the third position of the model string. | |
| − | + | model_list = re.findall("'[Z0-9] [Z0-9] Z [Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9]", stuff, re.M | re.S) | |
| + | Rerun ''btsummary.py'', and now the total matches should equal the number of model strings sampled in which q21=0. | ||
| + | <div style="background-color:#ccccff"> | ||
| + | * ''So what is the estimated marginal posterior probability that q21=0?'' {{title|I got 0.995|answer}} | ||
| + | * ''Why is the term marginal appropriate here (as in marginal posterior probability)?'' {{title|We are estimating the sum of all joint posteriors in which q21 equals 0|answer}} | ||
| + | </div> | ||
| + | |||
| + | == Estimating ancestral states == | ||
| + | |||
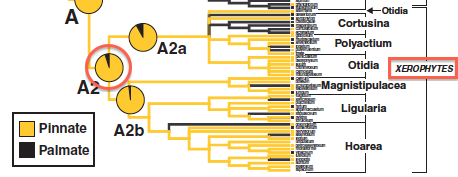
| + | [[File:Xerophytevenation.png|right]] The Jones et al. 2009 study estimated ancestral states using SIMMAP. In particular, they found that the most recent common ancestor (MRCA) of the xerophytic (dry-adapted) clade of pelargoniums almost certainly had pinnate venation (see red circle in figure on right). Let's see what BayesTraits says. | ||
| + | |||
| + | Start BayesTraits in the usual way, specifying 1 (Multistate) on the first screen and 2 (MCMC) on the second. After the options are output, type the following commands in, one line at a time, finishing with the run command: | ||
| + | pa exp 30 | ||
| + | addtag xerotag alternans104 rapaceum130 | ||
| + | addmrca xero xerotag | ||
run | run | ||
| − | The | + | The addmrca command tells BayesTraits to add columns of numbers to the output that display the probabilities of each state for each character in the most recent common ancestor of the taxa listed in the addtag command (2 taxa are sufficient to define the MRCA, but more taxa may be included). The column headers for the last four columns of output should be (I've added the comments starting with <--) |
| + | xero - S(0) - P(0) <-- character 0 (dissection), probability of state 0 (unlobed) | ||
| + | xero - S(0) - P(1) <-- character 0 (dissection), probability of state 1 (dissected) | ||
| + | xero - S(1) - P(0) <-- character 1 (venation), probability of state 0 (pinnate) | ||
| + | xero - S(1) - P(1) <-- character 1 (venation), probability of state 1 (palmate) | ||
| + | |||
| + | You can download the output file and view it in Tracer. That way you can use Tracer to tell you the means of the four columns above. Note that you will need to remove the initial text from the file (but keep the column headers) before Tracer will recognize it. | ||
| + | |||
| + | <div style="background-color:#ccccff"> | ||
| + | * ''Which state is most common at the xerophyte MRCA node for leaf venation?'' {{title|pinnate venation; xero - S(1) - P(0)|answer}} | ||
| + | * ''Which state is most common at the xerophyte MRCA node for leaf dissection?'' {{title|dissected; xero - S(0) - P(1)|answer}} | ||
| + | </div> | ||
| + | |||
| + | That concludes the introduction to BayesTraits. A glance through the manual will convince you that there is much more to this program than we have time to cover in a single lab period, but you should know enough now to explore the rest on your own if you need these features. | ||
| − | ==== | + | == Challenge == |
| − | + | ||
| − | + | BayesTraits allows you to ''fossilize'' (or ''fossilise'' in British English) traits for specific nodes. The relevant section of the BayesTraits manual is titled ''Fixing node values / fossilising'' and, for this challenge, you will need to locate the table at the end of this section (just before the section titled ''Discrete'' begins). | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | Estimate the marginal likelihood under these 2 models, both using the dependence model and MCMC: | |
| + | * fossilize the MRCA of the xerophytic (dry-adapted) clade to have pinnate venation | ||
| + | * fossilize the MRCA of the xerophytic (dry-adapted) clade to have palmate venation | ||
| − | + | Note: the <tt>Node01</tt> in the manual is just a name you invent to identify this fossilization constraint; you could call this <tt>xeronode</tt> if you want. | |
| − | The | + | The main question is: What is the log Bayes Factor for pinnate vs. palmate venation? |
| − | The | + | Turn in the following: |
| + | * The log marginal likelihoods for the two models | ||
| + | * The commands you used to achieve these log marginal likelihoods in BayesTraits | ||
| + | * The log Bayes Factor you calculated | ||
| + | * Assuming the two models compared have equal prior probabilities, how many times more probable is pinnate than palmate leaves in the ancestor of the xerophyte clade? | ||
| − | + | Hint: for the last item, use e^{log BF} to convert your log Bayes Factor to a ratio of marginal likelihoods, then multiply by the ratio of model prior probabilities. This uses Bayes Rule to convert the likelihood of the model to the posterior probability of the model. | |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | |||
| − | |||
| − | |||
[[Category: Phylogenetics]] | [[Category: Phylogenetics]] | ||
Latest revision as of 15:20, 30 March 2020

|
EEB 349: Phylogenetics |
| In this lab you will learn how to use the program BayesTraits, written by Andrew Meade and Mark Pagel. BayesTraits can perform several analyses related to evaluating evolutionary correlation and ancestral state reconstruction in discrete morphological traits. |
Contents
Download BayesTraits
Login to Xanadu and request a machine as usual:
srun --pty -p mcbstudent --qos=mcbstudent bash
Download BayesTraits from Mark Pagel's web site using curl. You can get the tar archive linked to the web site as "BayesTraits V3.0.2 - Linux 64" onto Xanadu however you like, but I think the easiest way is to just use curl:
curl -O http://www.evolution.rdg.ac.uk/BayesTraitsV3.0.2/Files/BayesTraitsV3.0.2-Linux.tar.gz
Now unpack the gzipped "tape archive" as follows:
tar zxvf BayesTraitsV3.0.2-Linux.tar.gz
This will create a directory named BayesTraitsV3.0.2-Linux. The BayesTraitsV3.0.2-Linux folder contains the program itself along with several tree and data files (e.g. Primates.txt and Primates.trees). I will hereafter refer to the folder containing these files as simply the BayesTraits folder. Go back to Mark Pagel's web site and download the manual for BayesTraits. This is a PDF file and should open in your browser window.
A little aside on tar files
Data used to be stored on magnetic tape, not hard drives, and the tar (tape archive) program is what was used to move files to and from the tape. This tells you something about how old the tar format is because perhaps none of you have even seen a magnetic tape used for data storage! The tar command takes all the files in a directory and simply concatenates them into one gigantic file. It also preserves file permissions and the directory structure. The four letters after the command name tar are zxvf. These stand for the following:
- z = uncompress (the gz at the end of the file tells you it is a compressed archive, so the z tells tar to uncompress it before unpacking it)
- x = extract (unpack the archive into individual files. You would use c here if you were creating a tar file)
- v = verbose (tell us what's going on as you unpack)
- f = file (this tells tar that the file name is coming next, so don't put f earlier in the list)
This tar file has been compressed using the program gzip, which adds the gz ending to the file name. Most tar files are compresses with gzip or some similar algorithm so that the file requires less time to move across the internet.
Download the tree and data files
For this exercise, you will use data and trees used in the SIMMAP analyses presented in this paper (you should recognize the names of at least two of the authors of this paper):
Jones C.S., Bakker F.T., Schlichting C.D., Nicotra A.B. 2009. Leaf shape evolution in the South African genus Pelargonium L'Her. (Geraniaceae). Evolution. 63:479–497.
The data and trees were not made available in the online supplementary materials for this paper, but I have obtained permission to use them for this laboratory exercise.
- pelly.txt This is the data file. It contains data for two traits (see below) for 154 taxa in the plant genus Pelargonium.
- pelly.tre This is the tree file. It contains 99 trees sampled from an MCMC analysis of DNA sequences.
You should move these files to a new folder that you create for this lab. For example
cd # cd alone returns you to your home directory mkdir pelly cd pelly curl -O http://hydrodictyon.eeb.uconn.edu/people/plewis/courses/phylogenetics/data/pelly.txt curl -O http://hydrodictyon.eeb.uconn.edu/people/plewis/courses/phylogenetics/data/pelly.tre
Assessing the strength of association between two binary characters
The first thing we will do is see if the two characters (leaf dissection and leaf venation) in pelly.txt are evolutionarily correlated.
Trait 1: Leaf dissection
The leaf dissection trait comprises two states (I've merged some states in the original data matrix to produce just 2 states):
- 0 means leaves are entire (unlobed or shallowly lobed in the original study), and
- 1 means leaves are dissected (lobed, deeply lobed, or dissected in the original study).
Trait 2: Leaf venation
The leaf venation trait comprises two states:
- 0 means leaves are pinnately veined (one main vein runs down the long axis of the leaf blade), and
- 1 means leaves are palmately veined (several major veins meet at the base of the leaf).
To test whether these two traits are correlated, we will estimate the marginal likelihood under two models. The independence model assumes that the two traits are uncorrelated. The dependence model allows the two traits to be correlated in their evolution. The model with the higher marginal likelihood will be the preferred model. You will recall that we discussed both of these models in lecture, and also discussed the stepping-stone method that BayesTraits uses to evaluate models. You may wish to pull up those lectures to help answer the questions that you will encounter momentarily, as well as the BayesTraits manual.
Maximum Likelihood: Independence model
Type the following to start the BayesTraits program (assuming you are in the pelly folder and that the BayesTraitsV3.0.2-Linux is a "sister" folder:
../BayesTraitsV3.0.2-Linux/BayesTraitsV3 pelly.tre pelly.txt
You should see this selection appear:
Please select the model of evolution to use. 1) MultiState 2) Discrete: Independent 3) Discrete: Dependant 4) Continuous: Random Walk (Model A) 5) Continuous: Directional (Model B) 6) Continuous: Regression 7) Independent Contrast 8) Independent Contrast: Correlation 9) Independent Contrast: Regression 10) Discrete: Covarion
Press the 2 key and hit enter to select the Independent model. Now you should see these choices appear:
Please Select the analysis method to use. 1) Maximum Likelihood. 2) MCMC
Press the 1 key and hit enter to select maximum likelihood. Now you should see some output showing the choices you explicitly (or implicitly) made:
Options:
Model: Discete Independant
Tree File Name: pelly.tre
Data File Name: pelly.txt
Log File Name: pelly.txt.log.txt
Save Initial Trees: False
Save Trees: False
Summary: False
Seed 3162959925
Analsis Type: Maximum Likelihood
ML attempt per tree: 10
ML Max Evaluations: 20000
ML Tolerance: 0.000001
ML Algorithm: BOBYQA
Rate Range: 0.000000 - 100.000000
Precision: 64 bits
Cores: 1
No of Rates: 4
Base frequency (PI's) None
Character Symbols: 00,01,10,11
Using a covarion model: False
Restrictions:
alpha1 None
beta1 None
alpha2 None
beta2 None
Tree Information
Trees: 99
Taxa: 154
Sites: 1
States: 4
Now type run and hit enter to perform the analysis, which will consist of estimating the parameters of the independent model on each of the 99 trees contained in the pelly.tre file.
Tree No Lh alpha1 beta1 alpha2 beta2 Root - P(0,0) Root - P(0,1) Root - P(1,0) Root - P(1,1) 1 -157.362972 53.767527 34.523176 35.319157 20.707416 0.249998 0.250002 0.249998 0.250002 2 -158.179984 53.313539 34.182683 36.038859 20.997536 0.249999 0.250001 0.249999 0.250001 . . . 98 -156.647307 52.357626 36.749282 27.270771 13.086248 0.250244 0.249756 0.250244 0.249756 99 -156.532925 52.321467 36.641688 27.402067 13.200124 0.250234 0.249767 0.250233 0.249766
You will notice that BayesTraits created a new file: pelly.txt.Log.txt. Rename this file ml-independent.txt so that it will not be overwritten the next time you run BayesTraits:
mv pelly.txt.Log.txt ml-independent.txt
Try to answer these questions using the output you have generated (you'll need to consult the BayesTraits manual, but ask us if anything doesn't make sense):
- Which occurs at a faster rate: pinnate to palmate, or palmate to pinnate? answer
- Which occurs at a faster rate: entire to dissected, or dissected to entire? answer
- What do you think Root - P(1,1) means (i.e. the last column of numbers)? answer
Maximum Likelihood: Dependence model
Run BayesTraits again, this time typing 3 on the first screen to choose the dependence model and again typing 1 on the second screen to select maximum likelihood. You should see this output showing the options selected:
Options:
Model: Discete Dependent
Tree File Name: pelly.tre
Data File Name: pelly.txt
Log File Name: pelly.txt.log.txt
Summary: False
Seed 3601265953
Analsis Type: Maximum Likelihood
ML attempt per tree: 10
Precision: 64 bits
Cores: 1
No of Rates: 8
Base frequency (PI's) None
Character Symbols: 00,01,10,11
Using a covarion model: False
Restrictions:
q12 None
q13 None
q21 None
q24 None
q31 None
q34 None
q42 None
q43 None
Tree Information
Trees: 99
Taxa: 154
Sites: 1
States: 4
Run the analysis. Here is an example of the output produced after you type run to start the analysis. The column headers don't quite line up with the columns, but you can fix this in a text editor or by copying and pasting the table-like output from the log file into a spreadsheet program:
Tree No Lh q12 q13 q21 q24 q31 q34 q42 q43 Root - P(0,0) Root - P(0,1) Root - P(1,0) Root - P(1,1) 1 -151.930254 66.451053 37.783888 0.000000 62.220033 23.997490 23.299393 46.110432 36.632979 0.24999 0.249981 0.250026 0.250000 2 -152.925691 67.152271 38.611193 0.000000 60.925185 24.514488 23.937433 45.313366 37.199310 0.24999 0.249983 0.250023 0.250001 . . . 98 -150.816306 36.534843 27.359325 0.000000 66.563262 19.823546 24.944519 63.940577 31.074092 0.250048 0.249750 0.250304 0.249898 99 -150.712705 37.316351 27.260833 0.000000 64.364694 20.107653 25.004246 60.945163 31.658536 0.250030 0.249779 0.250272 0.249919
Before doing anything else, rename the file pelly.txt.Log.txt to ml-dependent.txt so that it will not be overwritten the next time you run BayesTraits.
Try to answer these questions using the output you have generated:
- What type of joint evolutionary transitions seem to often have very low rates (look for an abundance of zeros in a column)? answer
- What type of joint evolutionary transitions seem to often have very high rates (look for columns with rates in the hundreds)? answer
Bayesian MCMC: Dependence model
Run BayesTraits again, typing 3 on the first screen to choose the dependence model and this time typing 2 on the second screen to select MCMC. You should see this output showing the options selected:
Options:
Model: Discete Dependent
Tree File Name: pelly.tre
Data File Name: pelly.txt
Log File Name: pelly.txt.log.txt
Summary: False
Seed 3792635164
Precision: 64 bits
Cores: 1
Analysis Type: MCMC
Sample Period: 1000
Iterations: 1010000
Burn in: 10000
MCMC ML Start: False
Schedule File: pelly.txt.log.txt.Schedule.txt
Rate Dev: AutoTune
No of Rates: 8
Base frequency (PI's) None
Character Symbols: 00,01,10,11
Using a covarion model: False
Restrictions:
q12 None
q13 None
q21 None
q24 None
q31 None
q34 None
q42 None
q43 None
Prior Information:
Prior Categories: 100
q12 uniform 0.00 100.00
q13 uniform 0.00 100.00
q21 uniform 0.00 100.00
q24 uniform 0.00 100.00
q31 uniform 0.00 100.00
q34 uniform 0.00 100.00
q42 uniform 0.00 100.00
q43 uniform 0.00 100.00
Tree Information
Trees: 99
Taxa: 154
Sites: 1
States: 4
Before typing run type the following command, which tells BayesTraits to change all priors from the default Uniform(0,100) to an Exponential distribution with mean 30:
pa exp 30
- Why am I suggesting this switch? answer
- Why 30? answer
Also type the following to ask BayesTraits to perform a stepping-stone analysis:
stones 100 10000
Now run the analysis. This will estimate 100 ratios to brook the gap between posterior and prior, using a sample size of 10000 for each "stone". Here is an example of the output produced after you type run to start the analysis:
Iteration Lh Tree No q12 q13 q21 q24 q31 q34 q42 q43 Root - P(0,0) Root - P(0,1) Root - P(1,0) Root - P(1,1) 11000 -155.195365 78 14.423234 34.800270 8.845985 45.927148 12.622435 50.476188 52.844895 32.149168 0.250068 0.249969 0.249994 0.249968 12000 -154.161705 82 64.601017 12.382781 9.259134 51.796365 12.002095 23.744903 30.316089 21.865930 0.249936 0.249957 0.250095 0.250012 . . . 1009000 -154.343996 30 33.555198 50.086092 11.294490 38.518607 24.461032 47.295157 43.477964 21.726938 0.250057 0.249939 0.250045 0.249959 1010000 -154.195259 87 29.584898 35.410909 2.003582 61.981073 16.976124 14.895266 49.111354 14.419644 0.251115 0.247854 0.252551 0.248480
Before doing anything else, rename the file pelly.txt.Log.txt to mcmc-dependent.txt, and pelly.txt.log.Stones.txt to mcmc-dependent.Stones.txt so that they will not be overwritten the next time you run BayesTraits.
You will notice a column not present in the likelihood analysis named Tree No that shows which of the 99 trees in the supplied pelly.tre treefile was chosen at random to be used for that particular sample point. BayesTraits is sampling trees from the posterior distribution here; it cannot actually sample trees from the posterior because we have given it only data for two morphological characters, which would not provide nearly enough information to estimate the phylogeny for 154 taxa. It is as if we had given BayesTraits sequence data as well as our 2 morphological characters and it was using only the sequence data to estimate the posterior distribution of trees and edge lengths and only the morphological data to estimate rates for the morphological characters.
Try to answer these questions using the output you have generated:
- What is the log marginal likelihood estimated using the stepping-stone method? This value is listed on the last line of the file mcmc-dependent.Stones.txt (your value may differ from mine slightly) answer
Bayesian MCMC: Independence model
Run BayesTraits again, this time specifying the Independent model, and again using MCMC, pa exp 30, and stones 100 10000. Rename the output file from pelly.txt.log.txt to mcmc-independent.txt. Also rename pelly.txt.log.Stones.txt to mcmc-independent.Stones.txt.
- What is the estimated log marginal likelihood for this analysis using the stepping-stone method? answer
- Which is the better model (dependent or independent) according to these estimates of marginal likelihood? answer
Bayesian Reversible-jump MCMC
Run BayesTraits again, specifying Dependent model, MCMC and, this time, specify the reversible-jump approach using the command
rj exp 30
The previous command also sets the prior. Type run to start, then when it finishes rename the output file rjmcmc-dependent.txt.
The reversible-jump approach carries out an MCMC analysis in which the number of model parameters (the dimension of the model) potentially changes from one iteration to the next. The full model allows each of the 8 rate parameters to be estimated separately, while other models restrict the values of some rate parameters to equal the values of other rate parameters. The output contains a column titled Model string that summarizes the model in a string of 8 symbols corresponding to the 8 rate parameters q12, q13, q21, q24, q31, q34, q42, and q43. For example, the model string "'1 0 Z 0 1 1 0 2" sets q21 to zero (Z), q13=q24=q42 (parameter group 0), q12=q31=q34 (parameter group 1), and q43 has its own non-zero value distinct from parameter groups 0 and 1.
You could copy the "spreadsheet" part of the output file into Excel and sort by the model string column, but let's instead use Python to summarize the output file. Download (e.g. using curl) the file btsummary.py file and run it as follows:
python btsummary.py
This should produce counts of model strings. (If it doesn't, check to make sure your output file is named rjmcmc-dependent.txt because btsummary.py tries to open a file by that name.) Answer the following questions using the counts provided by btsummary.py.
- Which model string is most common? answer
- What does this model imply? answer
Notice that many (but not all) model strings have Z for q21. One way to estimate the marginal posterior probability of the hypothesis that q21=0 is to sum the counts for all model strings that have Z in that third position corresponding to q21. While it is pretty easy to add these numbers in your head, let's modify btsummary.py to do this for us (this might come in useful if you ever encounter results that are more complex): open btsummary.py and locate the line containing the regular expression search that pulls out all the model strings from the BayesTrait output file:
model_list = re.findall("'[Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9]", stuff, re.M | re.S)
The re.findall function performs a regular expression search of the text stored in the variable stuff looking for strings that have a series of 8 space-separated characters, each of which is either the character Z or a digit between 0 and 9 (inclusive). Copy this line, then comment out one copy by starting the line with the hash (#) character:
#model_list = re.findall("'[Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9]", stuff, re.M | re.S)
model_list = re.findall("'[Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9]", stuff, re.M | re.S)
Now modify the copy such that it counts only models with Z in the third position of the model string.
model_list = re.findall("'[Z0-9] [Z0-9] Z [Z0-9] [Z0-9] [Z0-9] [Z0-9] [Z0-9]", stuff, re.M | re.S)
Rerun btsummary.py, and now the total matches should equal the number of model strings sampled in which q21=0.
- So what is the estimated marginal posterior probability that q21=0? answer
- Why is the term marginal appropriate here (as in marginal posterior probability)? answer
Estimating ancestral states
The Jones et al. 2009 study estimated ancestral states using SIMMAP. In particular, they found that the most recent common ancestor (MRCA) of the xerophytic (dry-adapted) clade of pelargoniums almost certainly had pinnate venation (see red circle in figure on right). Let's see what BayesTraits says.Start BayesTraits in the usual way, specifying 1 (Multistate) on the first screen and 2 (MCMC) on the second. After the options are output, type the following commands in, one line at a time, finishing with the run command:
pa exp 30 addtag xerotag alternans104 rapaceum130 addmrca xero xerotag run
The addmrca command tells BayesTraits to add columns of numbers to the output that display the probabilities of each state for each character in the most recent common ancestor of the taxa listed in the addtag command (2 taxa are sufficient to define the MRCA, but more taxa may be included). The column headers for the last four columns of output should be (I've added the comments starting with <--)
xero - S(0) - P(0) <-- character 0 (dissection), probability of state 0 (unlobed) xero - S(0) - P(1) <-- character 0 (dissection), probability of state 1 (dissected) xero - S(1) - P(0) <-- character 1 (venation), probability of state 0 (pinnate) xero - S(1) - P(1) <-- character 1 (venation), probability of state 1 (palmate)
You can download the output file and view it in Tracer. That way you can use Tracer to tell you the means of the four columns above. Note that you will need to remove the initial text from the file (but keep the column headers) before Tracer will recognize it.
- Which state is most common at the xerophyte MRCA node for leaf venation? answer
- Which state is most common at the xerophyte MRCA node for leaf dissection? answer
That concludes the introduction to BayesTraits. A glance through the manual will convince you that there is much more to this program than we have time to cover in a single lab period, but you should know enough now to explore the rest on your own if you need these features.
Challenge
BayesTraits allows you to fossilize (or fossilise in British English) traits for specific nodes. The relevant section of the BayesTraits manual is titled Fixing node values / fossilising and, for this challenge, you will need to locate the table at the end of this section (just before the section titled Discrete begins).
Estimate the marginal likelihood under these 2 models, both using the dependence model and MCMC:
- fossilize the MRCA of the xerophytic (dry-adapted) clade to have pinnate venation
- fossilize the MRCA of the xerophytic (dry-adapted) clade to have palmate venation
Note: the Node01 in the manual is just a name you invent to identify this fossilization constraint; you could call this xeronode if you want.
The main question is: What is the log Bayes Factor for pinnate vs. palmate venation?
Turn in the following:
- The log marginal likelihoods for the two models
- The commands you used to achieve these log marginal likelihoods in BayesTraits
- The log Bayes Factor you calculated
- Assuming the two models compared have equal prior probabilities, how many times more probable is pinnate than palmate leaves in the ancestor of the xerophyte clade?
Hint: for the last item, use e^{log BF} to convert your log Bayes Factor to a ratio of marginal likelihoods, then multiply by the ratio of model prior probabilities. This uses Bayes Rule to convert the likelihood of the model to the posterior probability of the model.