Phylogenetics: r8s Lab

|
This article is still under construction. Expect it to change frequently until this notice is removed. |

|
EEB 349: Phylogenetics |
| The goals of this final lab are to learn how to: (1) set up a NEXUS file for the r8s program; (2) interpret the most important parts of the output of r8s; (4) view a tree with divergence times produced by r8s in TreeView. |
Using r8s to estimate divergence times
There is no version of r8s for the Windows operating system, so we will use the version installed on the Bioinformatics cluster instead.
Start the program PuTTY and connect to bbcxsrv1.biotech.uconn.edu
If you do not find PuTTY on your desktop, you can download the program again from the PuTTY website. Using PuTTY, connect to bbcxsrv1.biotech.uconn.edu.
Download r8s
r8s is installed on the cluster, so we don't need the program itself, but we do need the sample data file that comes with r8s for the lab today. I've placed the SAMPLE_SIMPLE file that comes with r8s on a web server, and you can download it directly to the cluster using the curl program. Note that you could download the program using a web browser, then transfer the file to the cluster using PSFTP (or some other SFTP program), but using curl is easier and faster:
First, create a directory for this lab in your home directory:
mkdir $HOME/r8slab cd $HOME/r8slab
Now, download the SAMPLE_SIMPLE data file there:
curl -o SAMPLE_SIMPLE http://hydrodictyon.eeb.uconn.edu/eeb349/SAMPLE_SIMPLE
Use the more command to see the first few lines of the SAMPLE_SIMPLE file. Note that r8s uses the NEXUS format for its input files. You can see more of the file by pressing the spacebar, and you can quit viewing the file by pressing the letter q (for quit).
See if you can answer these questions just by looking at the SAMPLE_SIMPLE file:
- Which NEXUS blocks are present? (Hint: you should find 2 NEXUS blocks in this file)
- Why does r8s need to know the number of sites? (as in the "blformat nsites =952 lengths=persite" command)
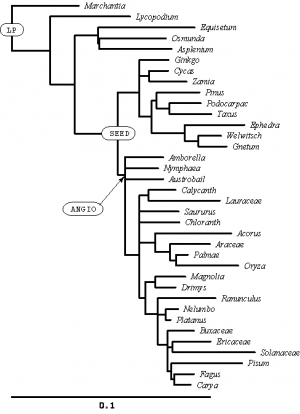
In the SAMPLE_SIMPLE file, these two lines define names for two interior nodes of the tree:
mrca LP marchantia pisum; mrca ANGIO amborella pisum;
I have labeled these nodes on the tree (right). Can you see how these simple mrca (Most Recent Common Ancestor) statements can uniquely define interior nodes?
Create an mrca statement to create a name for the node I have labeled SEED (there are many different ways you could do this, all of which would uniquely specify the labeled node).
Add your statement in the SAMPLE_SIMPLE file beneath the other two using the pico editor:
pico SAMPLE_SIMPLE
The command above will invoke the pico text editor program and you should see the contents of the file displayed in the main editor window. Use the arrow keys to navigate down to the two existing mrca lines, then add your mrca command after these. To exit pico, use the Ctrl-X key combination (the ^ symbol in the hints at the bottom of the window refers to the Ctrl key), then answer Y when asked if you want to save the modified buffer, and finally just press the Enter key when asked "File name to write : SAMPLE_SIMPLE"
- Which line in the data file serves to calibrate the (relaxed) clock?